REU Participating Faculty
- Noe Alvarez, analytical chemistry, nanomaterials
- Neil Ayres, polymer chemistry
- Ruxandra Dima, physical, computational chemistry (co-PI)
- Tony Grillo, biochemistry, bioanalytical chemistry
- Hairong Guan, inorganic & organic chemistry
- Anna Gudmundsdottir (PI), organic chemistry, material science
- Jianbing Jiang, Material science, catalysis, organic & inorganic chemistry
- In-Kwon Kim, biochemistry
- Wei Liu, organic synthesis, organometallic chemistry, and chemical catalysis
- James Mack, organic chemistry, green chemistry
- Ashley Ross, analytical chemistry, neuroscience
- George Stan, physical, computational chemistry
- Pietro Strobbia, analytical chemistry, spectroscopy and nanotechnology
- Yujie Sun, inorganic, catalysts
- Ryan White, analytical chemistry
Research Projects
The research in our group is focused on carbon nanomaterials synthesis and assembly into macroscopic materials for sensor applications. We synthesize and assemble carbon nanotubes into fibers and films and use them for physiological and electrochemical sensors, as well as energy storage devices. Besides of the fundamental chemistry such as synthesis and electrochemistry, students in the Lab are exposed to engineering aspects of nanomaterials development. We are developing microelectrodes suitable for physiological applications such as recording extracellular activity for neuroscience research and the ability to stimulate neurons in a targeted fashion. This research is focused on a bottom-up approach that allows us to combine carbon nanotubes (CNTs) into macroscopic flexible electrodes that can be adjusted to application-specific requirements. Electron transfer rates are currently under study using Electroretinogram (ERG) for signal recording and others for electrical stimulation. Biocompatible polymer coatings and control over their porosity and stiffness are topics of interest in neuroscience as implants to prevent damaging brain tissue. Additionally, we work on heavy metals detection in drinking water that has become high priority for our society, particularly for people living in cities where water infrastructure was built more than 30 years ago. This research intents to develop an electrochemical sensor based on CNTs to detect toxic metals such as Pb, Cd and Hg and can monitor them autonomously.
The use of polymers in society has become ubiquitous, from commodity items to high technology and cutting-edge therapeutics and diagnostics. The research in our group is focused on synthetic chemistry approaches to studying new biomaterials. Specifically, we are interested in blood-contacting biomaterials and co-polymer foams. To accomplish our goals, we use polymer chemistry to mimic naturally occurring biological macromolecules. In addition to uses as new biomaterials, the polymers prepared in our group can be used as shape-memory polymers, self-healing materials, gel-forming materials, and rheology modifiers. REU students in our group will contribute to using this chemistry to prepare polymers designed for determining the structure/activity relationships of synthetic heparin mimics. This project will result in a rationally designed set of materials with precise architecture and functional groups that will provide new, fundamental insights into polymer/host interactions. Students involved in this project will gain experience in polymer synthesis and characterization techniques, experimental design, and problem solving techniques.
Research in the Dima group focuses on understanding the role of various structural and cellular factors in the mechanical response of biological molecules ranging from RNA molecules and multi-domain proteins to large fibrillar assemblies that play crucial roles in fundamental processes such as the maintenance of the cell shape, cell mobility, cell-cell adhesion, axonal growth, and cellular division (mitosis). We use a variety of computational methods such as molecular simulations, bioinformatics, and machine learning.
A project for an REU student is “Exploring the dynamics of protein complexes using modeling and machine learning". Microtubules, large multi-filament polymeric complexes which are the main component of the cell cytoskeleton, play fundamental roles in cellular processes ranging from cellular transport to mitosis. These roles are all intimately connected with microtubules' ability to depolymerize under controlled cellular conditions. This control is exerted by a large array of microtubule associated proteins (300 or so species), which form transitory complexes with microtubules over different timescales. Recent experimental results strongly suggest that many of these protein co-factors are molecular machines that work by converting chemical energy into mechanical work, which is then applied to the microtubule polymer lattice. However, little is known about the details of the process. The goal of this project is to determine the relationship between the steps in the functional cycle of such molecular machines and the structural changes taking place inside the protein monomers of the machines and of their substrates, which are important during processes such as cell division or cell-cell interactions.
A student involved in this project will become familiar with (1) the computational methodology behind realistic modeling of proteins and their cellular environment (water, ions), (2) machine learning (data science) methods in biochemistry, (3) data analysis that couples results of simulations with experimentally derived data, (4) how to use visualization tools for large biomolecules, (5) how to search and analyze protein databases, and (6) how to search the scientific literature.
Age is the greatest risk factor in acquiring most mortality-causing diseases such as Alzheimer's disease, heart disease, diabetes, and cancer. Mitochondrial dysfunction remains one of the earliest features in these diseases. However, there remains an urgent need to better understand the influence of mitochondrial dysfunction on the molecular determinants that elicit aberrant metabolism. Our lab probes the implications of mitochondrial dysfunction in protein toxicity, aggregation, and misregulation to illuminate the underpinnings of mitochondrial dysfunction in neurodegenerative diseases. To achieve this, we use a variety of biochemical, cellular, and animal models with the goal of discovering novel pharmaceutical interventions to treat biologic decline with age. An REU student would learn common biochemical or molecular biology techniques to isolate biomolecules and analyze protein or RNA expression. They will use these methods to determine the changes in protein regulation in models of neurometabolic diseases.
The research in our group lies at the interface between inorganic and organic chemistry, focusing on the development of homogeneous catalysts based on first-row transition metals such as nickel, cobalt, iron, and copper. Such efforts are motivated by the fact that precious metals, which are widely used today in catalysis for synthesizing commodity and specialty chemicals, are expensive, limited in supply, and sometimes difficult to remove from organic products. The challenge of using first-row transition metals for catalysis starts with the difficulty in identifying ligands that can not only bind tightly to the metals but also promote precious metal-like reactivity or de-emphasize the metal’s role. Our investigation of pincer-ligated metal hydrides has led to the discovery of nickel- and iron-based catalysts for the hydroboration of CO2 to methanol derivatives and the hydrogenation of fatty acid methyl esters to fatty alcohols. Our ongoing projects build on these initial successes and focus on the improvement of catalytic efficiencies through further modification of the catalyst structures. REU students involved in these projects will learn various synthetic techniques, including the handling of air- and moisture-sensitive compounds. They will also be trained to conduct mechanistic studies using NMR spectroscopy, X-ray crystallography, and chemical kinetics. Furthermore, the research projects will teach students the concept of increasing energy efficiency by performing catalytic reactions and the notion of sustainability by using renewable feedstock and readily available materials.
The quest for new smart organic materials capable of transducing light and other stimuli into mechanical motion has intensified in recent years because of their potential in various applications, including biomedicine and soft robotics. We are investigating release of nitrogen molecules from crystalline organic azido derivatives. Upon exposure to light azido crystals respond very differently, some of them dance around, other crack and fracture fiercely, whereas others gently coil. We are identifying the reaction mechanism for various azido derivatives and correlating the rate of release of the nitrogen molecules with the crystal packing of the starting material. An REU student would spend the summer being trained to synthesize new azido derivatives and study the mechanism of their photoreactivity using transient spectroscopy and product studies, and document the crystal movements. The proposed research would allow the student to become familiar with using various spectroscopic methods, such as NMR, IR, and UV absorption.
Renewable Energy Projects in the Jiang Group
Our research interests center on the development of advanced materials with novel properties to address the main question in the renewable energy field. Leveraging synthetic and molecular engineering techniques, we will design and synthesize a series of model materials to understand the molecular interactions underpinning renewable energy storage. Our research is focused in two areas: catalysis for renewable fuel production and rechargeable battery for energy storage, as briefly described below.
Project 1. Redox Flow Battery for Large-Scale Energy Storage Applications
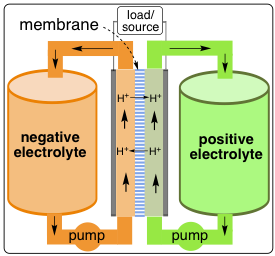
Fig 1. RFB Configuration
Among several available rechargeable battery technologies, redox flow batteries (RFBs) have received increasing attention as a safe and cost-effective energy storage technology for stationary energy storage application due to their safety, cost efficiency and scalability. Compared to traditional batteries in which redox-active materials are confined to the solid-state electrodes, the decoupled control of energy and power of RFBs enables an independent control of capacity and power to meet different requirements. Increasing the amount of energy to be stored involves simply using larger amounts of electrolytes (larger reservoirs, Fig 1), while increased power can be achieved by increasing the surface area of the electrodes and membrane in the current collector. While great progress has been made in the past a few years, three major issues remain: (1) low volumetric energy density. (2) High price, mainly due to the use of expensive Nafion membranes, and (3) The crossover of electrochemical active species from one electrolyte to the other lowers the overall Coulombic efficiency, and causes long-term, irreversible contaminations of the electrolytes. Very few current RFB systems possess the necessary combination of high electrolyte solubility, multielectron activity, and low cost to be viable for practical applications. Our research interest focuses on the design, synthesis and characterization of multielectron-active, highly soluble, polymeric materials with tunable redox properties as both anolyte and catholyte for RFBs applications that overcome the above-mentioned challenges. We developed a strategy of incorporating poly(ethylene glycol) units onto the energy storage materials to substantially improve the overall battery performance. The research outcome will be novel organic materials with unprecedented multielectron-activity, alleviated crossover, and improved energy density for practical applications.
Project 2. Efficient CO2-to-CO conversion by a Zinc Porphyrin Complex with a Triazole Unit
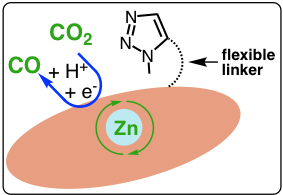
Fig 2, Catalytic CO2 conversion
One of the most attractive approaches towards providing carbon-neutral energy is the electrochemical conversion of CO2 from fossil fuel burning back to fuel or chemical feedstocks. Since the chemical reduction of CO2 is energetically uphill due to its remarkable thermodynamic stability and high activation energy associated with the multi-electron transfer steps, this process requires a significant amount of energy. It is imperative to apply efficient electrocatalysts to accelerate the sluggish reaction kinetics and to improve product distribution. Despite the notable progress, it still remains a great challenge to steer the reaction pathways toward desirable products and to reduce reaction overpotentials, necessitating the development of better electrocatalysts with higher product selectivity and catalytic activity. Our group designed, synthesized and tested a zinc-porphyrin complex with a flexible secondary coordinating unit triazole for efficient CO2 conversion to CO in 99% Faradaic efficiency. The triazole-free analogue gives lower catalytic current density with lower Faradaic efficiency. This work represents the first study of using a flexible triazole unit to enable CO2 reduction by a molecular zinc complex. We hypothesize that the triazole unit functions as a proton delivery site to facilitate proton shift to the catalytic site for CO2 conversion. Spectroscopy, voltammetry and isotopic effect techniques will be employed to investigate catalytic processes to gain mechanistic insights.
Regulation of poly(ADP-ribosy)lation (PARylation) by poly(ADP-ribose) turnover, a reversible post-translational modification, is essential for cellular responses to DNA damage and cell fate determination. Small-molecule inhibitors targeting this pathway has shown promise in specifically killing breast cancers. We are highly interested in determining the crystal structures of and developing a high-throughput assay for poly(ADP-ribose) turnover proteins. These are the pre-requisite steps to discover new cancer therapeutics through a high-throughput screening. A REU student will be primarily trained in expression, purification, and crystallization of proteins to determine the structures of proteins, protein-ligand, and protein-protein complexes.
The Liu research group focuses on the development and understanding of transition metal-catalyzed bond-forming reactions. The scope of our studies encompasses the synthesis and characterization of putative organometallic intermediates, the synthesis of biologically relevant molecules, and the creation of new activation modes for transition metal catalysis. These include the synthesis of high-valent copper(III) complexes that can undergo C–C bond-forming reductive elimination reactions, copper-catalyzed difluoromethylation reactions, and aryl radical activation strategies for cross-couplings. The students working on these projects will gain experience in organic synthesis, organometallic chemistry, and chemical catalysis.
The research focus of our group is the development and understanding of solvent-free organic reactions. In our research laboratory, we employ the novel solvent-free technique of mechanochemistry. Under this process, we use a high-velocity ball bearing to pulverize particles to the point that a reaction occurs. In addition to conducting solventless reactions, we also design our reactions such that they are environmentally friendly. Our purification procedure tries to minimize the use of harmful solvents, both in the reaction as well as at the purification stage. To create a new generation of environmentally conscious chemists that think of the environmental ramifications alongside the potential solutions to scientific problems, aspects of green chemistry must be taught early in their careers. REU students in our laboratory will work on various reactions using mechanochemistry and learn various aspects of green chemistry.Given the novelty of the technique, there are opportunities for the results of an REU student to be published in prominent scientific journals and presented at scientific meetings. Betsegaw Lemma, an NSF REU student in my research group recently co-authored a paper in Green Chemistry based on the research he conducted as an REU student in my lab. This research began a fruitful collaboration with Prof. Michael Coleman, Betsgaw’s undergraduate advisor. Since then, Professor Coleman, two more recent REU students, Sara Pilson and Devonna Leslie, and I published our latest collaborative project in Angew. Chem.7 We are currently in the process of submitting another manuscript based on their REU research.10 Many of my research papers have undergraduate students as co-authors, and I am a firm believer in incorporating undergraduate students fully into my research projects.
Our research group works in developing sensors and sensing platforms to study the brain and immune system. We are interested in unraveling the mechanisms of neurotransmitter regulated immunity in the contexts of health, autoimmunity, and neuroinflammation. Neurotransmitters are important chemical messengers in the brain and periphery which regulate a vast array of signaling transduction pathways; however, their unique role in regulating inflammatory responses is not well-understood. We use a combination of techniques including electrochemistry, microfluidics, and fluorescence microscopy to achieve these goals. Fast-scan cyclic voltammetry (FSCV) with carbon-fiber microelectrodes is an electrochemical technique that allows subsecond detection of electroactive neurotransmitters like dopamine, serotonin, and others in spatially discrete regions of tissue. Specific projects in our lab range from developing novel chemical sensors, fundamental investigations into analyte interactions at the electrode surface, developing microfluidic platforms for manipulating tissue and cells, to characterizing neurotransmitter signaling in vitro and ex vivo. REU students in our lab would learn how to fabricate and modify the surface of carbon-fiber microelectrodes, design and fabricate microfluidic devices using photolithography, and learn electrode surface analysis techniques.
Research in our group is focused on computational modeling of biological nanomachines involved in protein degradation. Protein quality control, such as degradation mechanisms, prevents deleterious off-pathway reactions of misfolding or aggregation. In the degradation pathway, AAA+ (ATPases associated with various cellular activities) nanomachines, such as bacterial caseinolytic protease (Clp) ATPases, unfold and translocate substrate proteins (SPs) through narrow central pores as a prerequisite for the ultimate destruction of the polypeptide chain within the peptidase chamber. Understanding the protein remodeling mechanisms of these biological nanomachines is of central importance for deciphering the details of cellular processes. A specific project for an REU student will concern probing the dependence of the Clp ATPase unfoldase function on the direction of applied mechanical force. Multiscale modeling of Clp-mediated unfolding of SPs with discernible mechanical anisotropy will yield detailed information of these properties. We propose to probe mechanisms of unfolding and translocation along the restricted direction of the N–C termini. The SPs to be studied will include wild-type and variants of proteins probed in single-molecule experiments. Unfolding and translocation pathways obtained in these studies will be contrasted with those in multidirectional pulling geometries, which mimic the cellular environment. Interactions between the Clp ATPase and SPs will probe multiple orientations of the substrate. The student working on this project will gain knowledge concerning biomolecular simulations of nanomachines and use of computational tools for visualization and analysis of biomolecular structures.
Our research group is interested in the development of optical sensing technologies for application in clinical settings or in the field. REU students in our program will be involved in the development and application of surface-enhanced Raman scattering (SERS) biosensors. SERS is a powerful type of spectroscopy that offers highly multiplexed and accurate detection of target molecules; however, SERS has not yet been utilized in real-world applications. We develop nanoparticles, sensing mechanism and devices to exploit these advantages in critical applications. We are developing optical SERS sensors to detect viral biomarkers at the point of care, as well as to detect dangerous contaminants directly in water. This research experience represents a multidisciplinary experience in analytical chemistry, spectroscopy and nanotechnology. REU students will learn nanoparticle synthesis and characterization, working with buffers and other elements used in bioanalysis, use of different optical-spectroscopy techniques and advanced data analysis methods.
The growing global energy demands, depletion of fossil fuel reserves, and increasing concerns about climate change resulting from fossil fuel utilization have urged the exploration of green and sustainable energy resources. Research in the Sun group spans the core disciplines of inorganic chemistry, materials science, electrochemistry, and photochemistry, focusing on the fundamental design, synthesis, and understanding of novel materials and complexes for renewable energy catalysis and sustainability. Currently, we are interested in a variety of energy-related reactions, such as electrocatalytic H2 oxidation and N2 reduction, as well as electrocatalytic and photocatalytic organic transformation. In order to gain the fundamental understanding of catalytic mechanisms of various reactions, we are devoted to designing and developing solid-state and molecular catalysts with well-defined active sites, ranging from metal/compound interfaces, ultrathin 2D nanosheets, single-atom catalysts, to heterobimetallic coordination complexes. REU students in our group will gain experience in the syntheses of solid-state materials and inorganic coordination complexes, as well as a variety of analytical and physical characterization techniques, such as NMR, MS, GC, HPLC, SEM, TEM, XRD, and XPS. Undergraduate students will also be trained in conducting electrochemical studies using cyclic voltammetry, linear sweep voltammetry, chronoamperometry, chronopotentiometry, Tafel analysis, rotating disk electrode voltammetry, etc. Our group has a strong track record in supporting undergraduate research projects and many of those undergraduate students have been co-authors on our publications.
Our research group is interested in the development of new electrochemical sensing and imaging methods to study biological systems with a high level of specificity and spatiotemporal resolution. REU students in our program will be involved, specifically, in the development and application of a class of electrochemical, aptamer-based sensors. More specifically, students will be exposed to senor fabrication and characterization. Unfortunately, there are no good methods for studying the thermodynamics of small molecule binding to an aptamer immobilized on an electrode surface. With their new knowledge of how to work with aptamer-based sensors, the student will use a newly described method of intermittent pulse amperometry to measure surface binding kinetics of small molecule targets to an electrode bound aptamer. By collecting kinetic time traces of association and dissociation processes, the student will fit the data to first-order reaction kinetics to learn more about the thermodynamics of the interaction between the small molecule targets and the aptamer-modified electrode. This research experience represents a multidisciplinary experience in analytical electrochemistry, biochemistry, and physical chemistry.